Traceability
Reproducibility asks "can I re-run this and get the same result?" Traceability asks a different but equally important question: "can I explain this result after the fact?" A traceable workflow makes it possible to look at any output and answer with confidence: which version of the code produced it, which input data was used, which parameters were set, and in what environment it ran. This is not just useful for debugging. In collaborative or regulated settings, it is essential for accountability. Think of traceability as the audit trail that sits behind reproducibility: one faces forward, the other looks back, and together they close the loop.
The mechanism that makes traceability possible on the code side is version control. Without a versioned codebase, you cannot reliably answer which code produced a given output, no matter how carefully you log everything else. But traceability does not stop at the code. The data that fed into a workflow, the parameters that shaped it, and the environment in which it ran all need to be captured too. This section works through each of those layers in turn.
Version control
The record
Version control is, at its most basic, a system that records the history of changes to a set of files. Every time you commit a change, you create a named, timestamped snapshot of the codebase. Every commit knows its parent. The full history of every file is preserved and queryable. This is what makes traceability on the code side possible: you can always travel back to any point in the history of a project and see exactly what the code looked like at that moment.
This has direct practical consequences for reproducibility. If a result was produced six months ago and a colleague asks which version of the pipeline generated it, the answer should be a commit hash, not a shrug. If a bug is discovered in a published analysis, the question of whether earlier results are affected is answerable from the version history, not from memory. If a workflow is updated and results change, the diff between two commits shows exactly what changed and why. None of this is possible without a versioned codebase.
It is also worth being explicit about something that often gets glossed over: version control is not the same as having backups or saving multiple copies of files with names like analysis_final_v3_revised.R. Those practices preserve files but they do not preserve history in a queryable, comparable, or reliable way. A proper version control system records not just what the code looks like now but the full sequence of intentional changes that brought it here, who made them, and when.
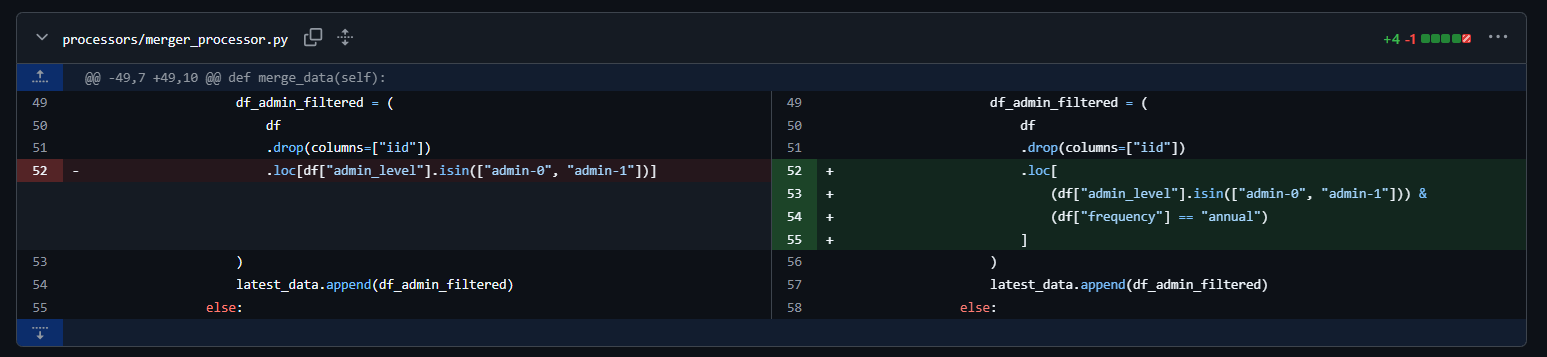
How does this looks in practice?
The discipline
Having Git initialized in a project is not the same as using version control well. The record is only as useful as the discipline behind it. A repository full of commits named "fix", "update", or "asdfgh" is technically versioned but practically untraceable. You cannot answer "which commit produced this result" if none of the commits describe what they did.
Good version control practice at the level of principles comes down to a few habits:
Commit messages should describe intent, not just action. "Fix null handling in income variable" is useful. "Fix bug" is not. A good commit message answers the question: if someone reads this in two years without any other context, will they understand what changed and why?
Branches isolate work in progress from stable code. Introducing a new analysis or testing a different model specification directly on the main branch means the stable version of the workflow is always at risk. Branches let you work in parallel without contaminating the main record. When the work is ready, it comes back through a merge or pull request.
Pull requests create a review checkpoint. Even on small teams, the practice of merging code through a reviewed pull request rather than pushing directly to the main branch creates a moment of deliberate scrutiny. It is the difference between a workflow that evolves through a series of intentional, reviewed decisions and one that accumulates changes informally.
Tags mark meaningful milestones. A commit hash is precise but not meaningful to a human reader. Tagging a commit (for example,
v1.0,paper-submission,baseline-model) creates a named, permanent reference to a specific state of the codebase that is both machine-readable and human-interpretable. For research workflows especially, tagging the exact version used to produce published results is a basic accountability practice.
None of this requires a complex branching strategy or a formal release process. The underlying principle is simply that changes to a workflow should be intentional, described, and recoverable. Version control is the system that enforces that standard.
Data versioning
Code versioning solves one half of the traceability problem. The other half is the data. A workflow is only fully traceable if you can answer not just "which code produced this output" but also "which version of the input data was used." Data changes too: sources get updated, files get corrected, pipelines get re-run on new extracts. Without some form of data versioning, two runs of the same code on nominally the same data can produce different results, and you may never know why.
Data hashing
The lightest-weight approach to data versioning is hashing. A cryptographic hash (typically SHA-256) is a fixed-length fingerprint of a file's contents. Change a single byte in the file and the hash changes completely. This makes it an extremely reliable way to verify that two copies of a file are identical, or to detect that a file has changed between runs.
In practice, storing a hash alongside every output is a minimal but powerful traceability guarantee. A JSON metadata file that records the SHA-256 hash of every input dataset used in a run means that anyone reproducing the workflow can verify that they are working with exactly the same data, or detect immediately if the source has changed.
Hashing is also the right tool when the data itself cannot be shared. If your workflow uses confidential, sensitive, or proprietary data, you cannot publish the data, but you can publish the hash. This gives other researchers working with the same dataset a way to verify that they have the right version, without requiring anyone to distribute the data itself. We touched on this in the Openness section in the context of sensitive data, and it is worth connecting the threads here: a hash is not a substitute for sharing data, but it is a meaningful reproducibility guarantee when sharing is not possible.
For hashing objects and files, two dependencies stand out as the natural choices for each language. In R, the digest package allows you to hash any R object directly (data frames, vectors, lists) without manual serialization, returning a hex string you can log or store. It supports multiple algorithms (MD5, SHA-1, SHA-256), is simple to use, and works on in-memory objects out of the box. The downside is that hashes of in-memory objects can be sensitive to things like column ordering or index state, which can make cross-environment comparisons fragile. In Python, the built-in hashlib module handles file-level hashing cleanly and is extremely stable across environments, since it operates on raw bytes. The trade-off is that hashing a DataFrame requires a manual serialization step first, typically via pd.util.hash_pandas_object(), which adds a bit of boilerplate. For both languages, hashing raw input files at the point of ingestion is generally more reproducible than hashing in-memory objects, and is the recommended approach for traceability purposes.
DVC: versioning data alongside code
Hashing is lightweight and universal, but it is manual. For workflows where data files are large, change frequently, or need to be shared across a team, a more structured approach becomes necessary. This is the problem that DVC (Data Version Control) was built to solve.
DVC extends Git to cover data and model artifacts. The core idea is that large files (datasets, model weights, processed outputs) are stored in a dedicated remote storage backend (an S3 bucket, Google Cloud Storage, a shared network drive, or any number of other options) while Git tracks a small, human-readable pointer file that records exactly which version of the data belongs with which version of the code. When you run dvc push, your data goes to the remote. When a collaborator runs dvc pull, they get exactly the version of the data that corresponds to the current Git commit.
# Track a data file with DVC
dvc add data/raw/survey.parquet
# This creates a .dvc pointer file that goes into Git
git add data/raw/survey.parquet.dvc .gitignore
git commit -m "Track raw survey data with DVC"
# Push data to remote storage
dvc pushThis connects back to something raised in the Openness section: the recommendation to always keep a copy of the data your workflow depends on, because external sources can disappear, change, or become inaccessible. DVC makes that practice systematic. Rather than manually managing copies of data files or relying on a URL that may stop working, DVC gives you a versioned, reproducible link between your code and your data that lives in your repository and is stored reliably in a backend you control.
For teams where multiple people are working on the same pipeline, or for workflows that run on scheduled jobs in different environments, the ability to dvc pull the exact right version of the data for any given Git commit is a meaningful traceability guarantee that hashing alone cannot provide.
DVC also integrates with experiment tracking, allowing you to version not just raw inputs but processed intermediates and model artifacts. For machine learning workflows in particular, where the same pipeline might be run dozens of times on different data slices or with different preprocessing steps, this level of data versioning quickly becomes indispensable.
If you want to learn more on DVC, it is worth knowing that they have a DVC Tools for Data Scientists & Analysts course on their website.
pins: A Lightweight Data Registry for R and Python
pins takes a different approach to data versioning than DVC. Rather than tying data history to a Git repository, it works around the concept of a board, a storage backend where you publish and retrieve versioned artifacts. Boards can be local folders, Posit Connect, Databricks, Amazon S3, Google Cloud Storage, Azure storage, or even Microsoft 365. The same mental model applies whether you are working in R or Python, and you can even use one language to read a pin created with the other.
The core workflow is simple. You write an artifact to a board with pin_write(), and retrieve it later with pin_read(). Each pin can be automatically versioned, making it straightforward to track changes, re-run analyses on historical data, and undo mistakes. Every version gets a timestamp-based identifier and a hash, so you can always retrieve a specific historical snapshot if needed.
In R:
library(pins)
board <- board_s3("my-bucket")
board |> pin_write(my_clean_data, "clean-survey-data", type = "parquet")
# retrieve it later
board |> pin_read("clean-survey-data")In Python:
from pins import board_s3
board = board_s3("my-bucket")
board.pin_write(my_clean_data, "clean-survey-data", type="parquet")
# retrieve it later
board.pin_read("clean-survey-data")Every pin is accompanied by metadata that includes a hash, file size, creation timestamp, and type, which makes it easy to log exactly what version of a dataset was used in a given run.
Where pins really shines is in team settings: a shared S3 bucket or a Posit Connect board lets collaborators publish and consume datasets without having to coordinate file transfers manually. It is also a natural fit for scheduled pipelines that need to hand off intermediate datasets between stages without relying on a shared filesystem.
That said, pins is not a replacement for DVC. It does not integrate with Git, it does not track how data changes in relation to code changes, and it is not appropriate when multiple sources or processes need to write to the same pin simultaneously, since it cannot manage concurrent writes. Think of it less as a version control system and more as a versioned data registry: a clean, low-friction way to publish, share, and retrieve artifacts with a traceable history. For projects that do not need the full DVC pipeline machinery, it is often the more practical choice.
Metadata capture at runtime
Version control tells you which code produced a result. Data versioning tells you which data was used. But there is a third layer of context that neither of those captures automatically: the runtime environment at the moment of execution. What parameters were passed? What timestamp? What version of the code was active? What input files were loaded?
The practice of capturing this as runtime metadata is simple but underused. The idea is to generate a small structured record alongside every meaningful output: a JSON sidecar file that sits next to your results table or model artifact and documents exactly the conditions under which it was produced. A minimal example in Python might look like this:
import json
import hashlib
import subprocess
from datetime import datetime, timezone
from pathlib import Path
def get_git_hash():
try:
return subprocess.check_output(
["git", "rev-parse", "HEAD"],
stderr=subprocess.DEVNULL
).decode().strip()
except Exception:
return "unavailable"
def hash_file(path: str) -> str:
h = hashlib.sha256()
with open(path, "rb") as f:
for chunk in iter(lambda: f.read(8192), b""):
h.update(chunk)
return h.hexdigest()
metadata = {
"timestamp": datetime.now(timezone.utc).isoformat(),
"git_commit": get_git_hash(),
"parameters": {
"model": "logistic_regression",
"test_size": 0.2,
"random_state": 42
},
"inputs": {
"training_data": {
"path": "data/processed/train.parquet",
"sha256": hash_file("data/processed/train.parquet")
}
}
}
Path("outputs/results_metadata.json").write_text(
json.dumps(metadata, indent = 2)
)This is deliberately minimal. The goal is not to build an elaborate logging system but to establish a habit: every output should have a record of what produced it. For more complex workflows, the tools covered in the next section handle much of this automatically.
Linear execution and the notebook trap
Before moving on to the tools that help with traceability, it is worth addressing one of the most common ways people accidentally undermine it: non-linear execution in notebooks.
Notebooks are genuinely useful tools. For exploring a dataset, prototyping an idea, or communicating a result with narrative and visuals, they are hard to beat. But they carry a reproducibility risk that is easy to underestimate: the ability to execute cells out of order, modify them after execution, and accumulate hidden state in memory. A notebook where cells have been run in arbitrary order may display results that cannot be reproduced by running it top to bottom. The code and the output are no longer telling the same story.
Pimentel et al. (2019) conducted large-scale studies on reproducibility issues across over 1.4 million Jupyter notebooks from GitHub, finding that only 24.11% of notebooks could be run without errors, and only around 4% produced the same results as originally reported. That is not a minor inconvenience. It means the vast majority of shared notebooks, even those attached to published work, cannot be independently verified.
The specific failure modes are well documented. Notebooks allow code cells to be executed out of order, leading to unexpected behaviour. For example, loading data in one cell, preprocessing it in subsequent cells, and training a model in another means that running those cells out of order can corrupt the entire pipeline. Similarly, cells can be modified after they have been executed, which can again lead to unexpected behaviour when the notebook is run from start to finish. This is what practitioners call hidden state: the kernel holds values in memory that are no longer reflected in the code as written, and the outputs become a product of the execution history rather than the code itself.
R softens this problem somewhat, particularly when working with Quarto documents, since rendering always executes the file from top to bottom in a fresh session. The rendered output is, by construction, a product of the code as written. But R is not immune to the hidden state trap, it just tends to show up elsewhere. The culprit is interactive execution in IDEs like RStudio or Positron, and it is surprisingly easy to fall into.
In my own experience supervising or working along junior analysts and interns, one pattern comes up repeatedly: R scripts run interactively, line by line, where the environment and the code quietly fall out of step. Changes made in the console may never reach the script, and changes made in the script may never reach the objects already sitting in memory that depend on them. The result is a script that appears to work but only produces the right output because of the specific sequence of interactive commands that preceded it. Running it top to bottom from a clean session gives a different answer, or fails entirely.
The rule is simple and non-negotiable: always verify that your script runs correctly from top to bottom in a clean environment before treating its output as final.
Personally, I have found that packaging my R code (more on this later) and/or creating the habit of running my R code from the terminal with Rscript my-script.R solves this issue, since it forces a clean session with no leftover environment state to hide your mistakes.
Finally, there is also a version control problem that is specific to Jupyter notebooks and worth calling out separately. Because .ipynb files are stored as JSON, they embed cell outputs, metadata, and execution counts alongside the code. A simple code change produces a messy, hard-to-read diff. Outputs committed to a repository can expose sensitive data. And merging changes from two contributors to the same notebook frequently results in corrupted files. This problem is considerably less severe with R Markdown (.Rmd) and Quarto (.qmd) files, which store only the source code and narrative in plain text, keeping diffs clean and outputs separate.
The practical guidance here is not "never use notebooks." It is closer to the principle that has become common in data engineering circles: use notebooks for exploration and communication, use scripts for production. Notebooks are the right tool when you are thinking out loud, interrogating data, or explaining something to an audience. They are the wrong tool when you need a process that someone else can run and get the same result.
Making that transition is not just about swapping file extensions. It is about adopting a different way of thinking about code organization altogether. As Lucy Dickinson describes in her guide The Journey from Jupyter to Programmer, the shift means moving toward modular, script-based workflows where code is broken into single-purpose functions, organized into clearly named files, and tied together through a single entry point. That structure is what unlocks the things that matter for reproducible work: proper version control, packaging, deployment, and the ability for someone else to run the whole thing from a clean environment without reverse-engineering your execution history. The notebook is still there for exploration. It just stops being the place where the final, trusted output lives.
INFO
A note on marimo
One of the fundamental complaints about Jupyter notebooks (the hidden state problem) is something that newer tools are beginning to address architecturally. Marimo is an open-source alternative to Jupyter notebooks where every cell is part of a dependency graph. Changes in one cell automatically trigger updates in dependent cells, keeping code and outputs in sync. Because a marimo notebook is stored as a pure Python file rather than JSON, it also integrates cleanly with version control, producing readable diffs and avoiding the output-leakage problems common in Jupyter. Marimo is a relatively recent tool and I have not used it extensively, but it is the most architecturally serious attempt to fix the reproducibility problems of traditional notebooks, and it is worth watching. I would suggest reading this blog post.
Other traceability tools
Python: MLflow
For Python workflows, MLflow has become the closest thing to a standard tool for experiment tracking and traceability. At its core, it provides a structured way to record the things that matter about a run: the parameters that were set, the metrics that were produced, the artifacts (models, plots, data files) that were generated, and the environment in which everything ran. All of this is stored in a local or remote tracking server and browsable through a web UI.
The basic workflow is straightforward. You wrap your code in an mlflow.start_run() context, log what you need, and MLflow handles the rest:
import mlflow
with mlflow.start_run():
mlflow.log_param("test_size", 0.2)
mlflow.log_param("random_state", 42)
mlflow.log_metric("accuracy", 0.87)
mlflow.log_metric("f1_score", 0.84)
mlflow.log_artifact("outputs/confusion_matrix.png")
mlflow.sklearn.log_model(model, "model")Beyond individual runs, MLflow makes it easy to compare experiments across runs, register model versions, and track which version of a model is in production versus staging. For teams working on multiple model iterations or running large parameter sweeps, this becomes genuinely indispensable.
MLflow is not limited to machine learning in the narrow sense. For any analytical workflow that involves parameters, metrics, and outputs worth tracking, it provides a cleaner and more queryable record than manual metadata files.
R: targets
For R, it is worth mentioning the targets package in the context of traceability, though it is not primarily a traceability tool. targets is a pipeline orchestration toolkit: you define your workflow as a set of named targets, each one a function call with declared inputs, and targets figures out which steps are up to date and which need to re-run when something changes. The traceability gains it provides, knowing what ran, what changed, and what produced what, are largely a byproduct of the dependency graph it builds to do its actual job.
Because its primary value is efficiency rather than accountability, we will cover targets in much more depth in the Optimality section, where pipeline orchestration and selective re-execution are the central topics. For now, it is enough to know it exists and that its self-documenting pipeline structure gives you a degree of traceability almost for free.