Portability
A workflow is portable when it runs correctly across different environments (your laptop, a colleague's machine, a cloud server) without requiring manual adjustments. The most common reason workflows break when moved is unmanaged dependencies: a package that is installed on your machine but not on someone else's, or a version that behaves differently across environments. This is the so-called "it-runs-in-my-laptop" problem.
There is a quick Stata example from the Reproducible Research Fundamentals course that illustrates how dangerous this can be. The idea is simple: create two random variables using different versions of the software and compare the results.
/* Stata */
set obs 10
version 13
set seed 1379
gen x1 = runiform()
set seed 1379
gen x2 = runiform()
version 14
set seed 1379
gen x3 = runiform()
br x1 x2 x3When we explore the resulting variables, we can see the following:
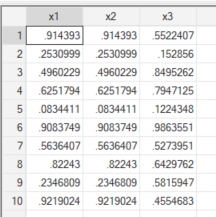
Because the underlying implementation of runiform() changed between versions, the two variables are different, even when we keep the same seeding. Same code, different results. This is a somewhat contained problem in Stata because most users stick to base commands, and declaring the Stata version at the top of a script is usually enough to pin behaviour. If you use ssc install commands, the safest practice is to save those packages inside a PLUS directory within your project so they are not pulled from whatever version happens to be installed on the current machine.
The R world has its own version of this story. If you have worked with R long enough, you have almost certainly encountered scripts still using dplyr's old scoped verbs: summarise_if(), mutate_at(), summarise_all(). All of these were deprecated in dplyr 1.0.0, released in 2020, in favour of across(). On a modern version of dplyr these calls will produce deprecation warnings; eventually, they will error entirely. The frustrating part is not the deprecation itself but the fact that those scripts rarely declare which version of dplyr they were written for. A bare library(dplyr) at the top of a file tells you nothing about the environment the author had in mind, and that is precisely the real problem. Using old versions of some libraries or deprecated functions is honestly fine, as long as we record the versions in which they are running in our workflows.
The solution to this problem is what it is known as environment isolation, and the right tool for that depends on how deep the isolation needs to go.
In day-to-day work, a virtual environment is almost always sufficient. A virtual environment isolates the language-level dependencies of a single project: the packages, the libraries, and the interpreter version. It lives entirely in user space, has near-zero overhead, and keeps project dependencies from interfering with each other. It is the tool you should reach for by default.
Occasionally, you will need something more. When a workflow depends not just on language packages but on system-level libraries — specific versions of things like libgdal, libpq, or a particular database client, a container becomes the right tool. A container (the most common implementation being Docker) isolates a full application environment, including OS libraries and system tools, while still sharing the host machine's kernel. It is heavier than a virtual environment but far more reproducible across machines with different operating system configurations. In practice, I reach for containers far less often than virtual environments, but when a pipeline has non-trivial system dependencies, they are hard to replace.
One honest caveat: containers can be difficult or impossible to use in certain institutional or corporate environments due to security and network restrictions. Docker traditionally requires root-level or equivalent privileges to run, which makes it a non-starter in many institutional environments where security policies restrict that kind of access. Rootless alternatives like Podman exist, but their availability depends entirely on your organization's infrastructure and policies. If that is your situation, documenting system dependencies carefully may be the only realistic path. Although, you could also try your luck and see if your institution would allow you to use Podman instead of Docker.
Python: uv
For Python projects, uv has become the tool I recommend without hesitation. To understand why, it helps to know what it replaces.
For a long time, managing a Python project meant juggling several separate tools: pyenv to manage Python versions, venv or virtualenv to create isolated environments, pip to install packages, and pip-tools to generate lockfiles that pin exact dependency versions. Each of these tools did its job reasonably well, but together they created a fragmented workflow with a lot of friction. uv replaces all of them. It manages Python versions, creates and manages virtual environments, resolves and installs dependencies, and generates lockfiles, all in a single tool, and extremely fast. The speed is not just a quality-of-life improvement: it changes how you interact with dependency management day to day. Running uv sync to restore an environment from a lockfile is fast enough that you stop dreading it.
The key files in a uv project are pyproject.toml, which declares your dependencies and project metadata, and uv.lock, which is the lockfile that pins every dependency to an exact version. Both should be committed to version control. When a collaborator clones your repository and runs uv sync, they get exactly the same environment you have.
INFO
Why uv and not conda?
If you learned Python in a scientific computing context, you probably learned conda first, and for good historical reasons. In the years before modern pip wheels and binary distributions became widespread, installing packages like NumPy or SciPy on Windows was genuinely painful. conda solved a real problem: it managed not just Python packages but also the system-level compiled libraries those packages depended on. For an entire generation of scientific Python developers, it was the only tool that made their environment actually work.
The problem is that conda solved that problem by creating a parallel ecosystem entirely separate from the standard Python toolchain. It introduced its own package repository (conda-forge), distinct from PyPI. It used its own environment and dependency model, incompatible with pip. Package maintainers who published to PyPI had to separately publish to conda-forge to reach conda users. And anyone who has worked on a mixed team of scientific developers (using conda) and software engineers (using pip and venv) has likely experienced the friction of those two ecosystems not playing well together, forcing painful migrations or fragile workarounds.
As pip matured and binary distributions became the norm, conda's core justification weakened considerably for most analytical workflows. It still has a legitimate use case: if your pipeline requires specific versions of system-level libraries like GDAL or HDF5, conda can manage those in a way that uv cannot. But for the vast majority of data and analytical work, that layer of complexity is not needed, and uv provides a cleaner, faster, and more interoperable alternative that keeps you firmly within the standard Python ecosystem.
For more information on this topic, I would suggest you to read this blog post from the Data Quarry
For a nice gentle introduction to how to use uv, I would suggest reading the following tutorial from DataCamp.
R: renv
For R, the renv package is the standard tool for project-level package management. The core workflow is straightforward: renv::init() sets up an isolated package library for your project, renv::snapshot() records the current state of that library into a lockfile (renv.lock), and renv::restore() reinstalls exactly those packages from the lockfile on another machine. The lockfile should always be committed to version control.
That said, the renv documentation is admirably honest about what the tool does not do, and it is worth repeating here. renv is not a panacea for reproducibility, it is a tool that helps with one part of the overall problem: R packages. Specifically, there are three things it does not handle:
- R version:
renvtracks, but does not help with, the version of R used with the project. For managing multiple R versions on one machine,rigis a practical companion tool. - Pandoc: the
rmarkdownpackage relies heavily on pandoc, but pandoc is not bundled with the package, meaning that restoringrmarkdownfrom the lockfile is insufficient to guarantee exactly the same rendering of RMarkdown documents. - System-level dependencies: operating system, versions of system libraries, and compiler versions are a separate challenge, but Docker is one popular solution. There is also a practical failure mode worth knowing about: package installation may fail if a package was originally installed through a binary, but that binary is no longer available.
renvwill attempt to install the package from source, but this can and often will fail due to missing system prerequisites.
None of this is a reason to avoid renv, it is still the right tool for R dependency management. But it is a reason to understand its boundaries and not treat a lockfile as a complete reproducibility guarantee on its own.
A note on nix and rix
For those who want to push portability further in R, it is worth knowing that the rix package offers a way to use the Nix package manager to declare fully reproducible environments that cover not just R packages but also the R version itself and system-level dependencies. This is a much stronger reproducibility guarantee than renv alone. Honest disclaimer: my experience with rix is limited, and it is still maturing as a tool. I would not recommend it as a starting point, but if you find yourself repeatedly running into the edges of what renv can do, it is a direction worth exploring.
Secrets and paths
Two smaller but important portability issues that come up constantly in practice.
Secrets and credentials (API keys, database passwords, authentication tokens) should never appear in your code or in any file committed to version control. The standard practice is to store them in environment variables and load them at runtime, typically via an external file that is explicitly excluded from version control through .gitignore. In Python, the python-dotenv package handles this cleanly, loading .env files. In R, the dotenv package or the built-in Sys.getenv() with an .Renviron file serve the same purpose. A workflow that embeds credentials in code is not just a security risk, it is a portability failure, because it will break the moment someone else tries to run it securely.
Hardcoded paths are the other common offender. A script that begins with something like /Users/JohnDoe/Documents/project/data/raw/file.csv will fail for every single person who is not you, and will likely fail for you too after a machine change or a folder reorganisation. Use relative paths, build paths programmatically from a project root, or use dedicated tools (here in R, pathlib in Python) that resolve paths relative to the project structure regardless of where it lives on disk.
Working with SharePoint
In many institutional settings, your organization will already have a file management platform in place, most commonly Microsoft SharePoint. This is not going away, and it doesn't need to. However, if you are not careful with how you are using both platforms and technoloies, you will be prono to conflicts in your code and syncing might start causing heavy headaches. The starting point is to be clear about what each of them is actually designed to do.
SharePoint is a document management and collaboration platform. Its versioning system tracks snapshots of files over time, it manages access permissions, and it gives non-technical colleagues a familiar interface to store and retrieve documents. For Word files, Excel spreadsheets, PDFs, and raw data files, it works perfectly well. What it cannot do is manage code. It has no concept of a commit message explaining why a change was made, no ability to show you a diff between two versions of a script, no branching, and no merging. SharePoint simply was not designed to manage code. It can store a script the same way it stores a Word document, but it has no understanding of what that file contains or how it has changed.
GitHub (or any Git-based platform) is a version control system for code. Every change is annotated with a message, an author, and a timestamp. You can branch off to work on a new feature without touching the main codebase, compare any two versions of a file line by line, and trace any piece of logic back to when and why it was introduced. None of this exists in SharePoint. They are solving genuinely different problems.
The practical recommendation here is to treat them accordingly: use SharePoint as a data lake from which you pull your inputs, and use GitHub as your day-to-day working platform. Raw data files, survey exports, administrative datasets, and any other inputs your organization stores centrally belong in SharePoint. Your code, your documentation, your project structure: all of that lives in GitHub, under version control, following the principles laid out in this guide.
If your organization also requires a copy of the codebase to be stored in SharePoint for compliance or visibility reasons, that is fine as a secondary archive. The important thing is that this copy is treated as a read-only snapshot, not a working copy. No collaborator should be pulling from it, editing files in it, or treating it as the source of truth for the project. The moment someone starts working out of the SharePoint copy instead of the GitHub repository, you have two diverging versions of the codebase and no good way to reconcile them.
On accessing data from SharePoint programmatically
There are solid options for both R and Python. In R, the {Microsoft365R} package provides a straightforward interface to SharePoint Online and OneDrive, authenticating through Azure Active Directory and allowing you to download files directly into your session without leaving a local copy on disk. In Python, Office365-REST-Python-Client covers the same ground. Both support interactive browser-based authentication, which is the safest default for day-to-day use.
That said, authentication can be a real friction point in practice. Registering an application in Azure AD (which enables the cleaner authentication flows) requires admin permissions that many data practitioners simply don't have. If that is your situation, the fallback is more manual: you sync the relevant SharePoint folder to your local machine using the OneDrive desktop client, and then reference the local synced path in your code.
This path will be different for every collaborator depending on their operating system and how they've set up their sync. The right way to handle this is to store it as an environment variable rather than hardcoding it. In R, this means adding a line like the following to your .Renviron file:
SHAREPOINT_DATA="C:/Users/yourname/YourOrg/ProjectName - Documents/data"And in Python, adding it to a .env file (loaded with a package like python-dotenv):
SHAREPOINT_DATA="/Users/yourname/YourOrg/ProjectName - Documents/data"Both .Renviron and .env files should be listed in your .gitignore so they never get committed to the repository. Each collaborator maintains their own local copy with their own path. The code itself then references the variable rather than any specific path:
# R
data_path <- Sys.getenv("SHAREPOINT_DATA")
df <- readr::read_csv(file.path(data_path, "raw", "survey_2024.csv"))# Python
import os
from dotenv import load_dotenv
import pandas as pd
load_dotenv()
data_path = os.environ["SHAREPOINT_DATA"]
df = pd.read_csv(os.path.join(data_path, "raw", "survey_2024.csv"))Whatever approach you end up using, make sure to document it clearly in your project's README. Collaborators joining the project should know immediately where the data lives, how to get access to it, and what environment variable they need to set before the code will run. This is not optional: it is one of the most basic requirements of a portable workflow.